The accuracy of protein structure prediction was evaluated using a Rosetta program. This repository contains an open-source protein distance prediction network, ProSPr, released under the MIT license. PROF: Secondary Structure Prediction System.
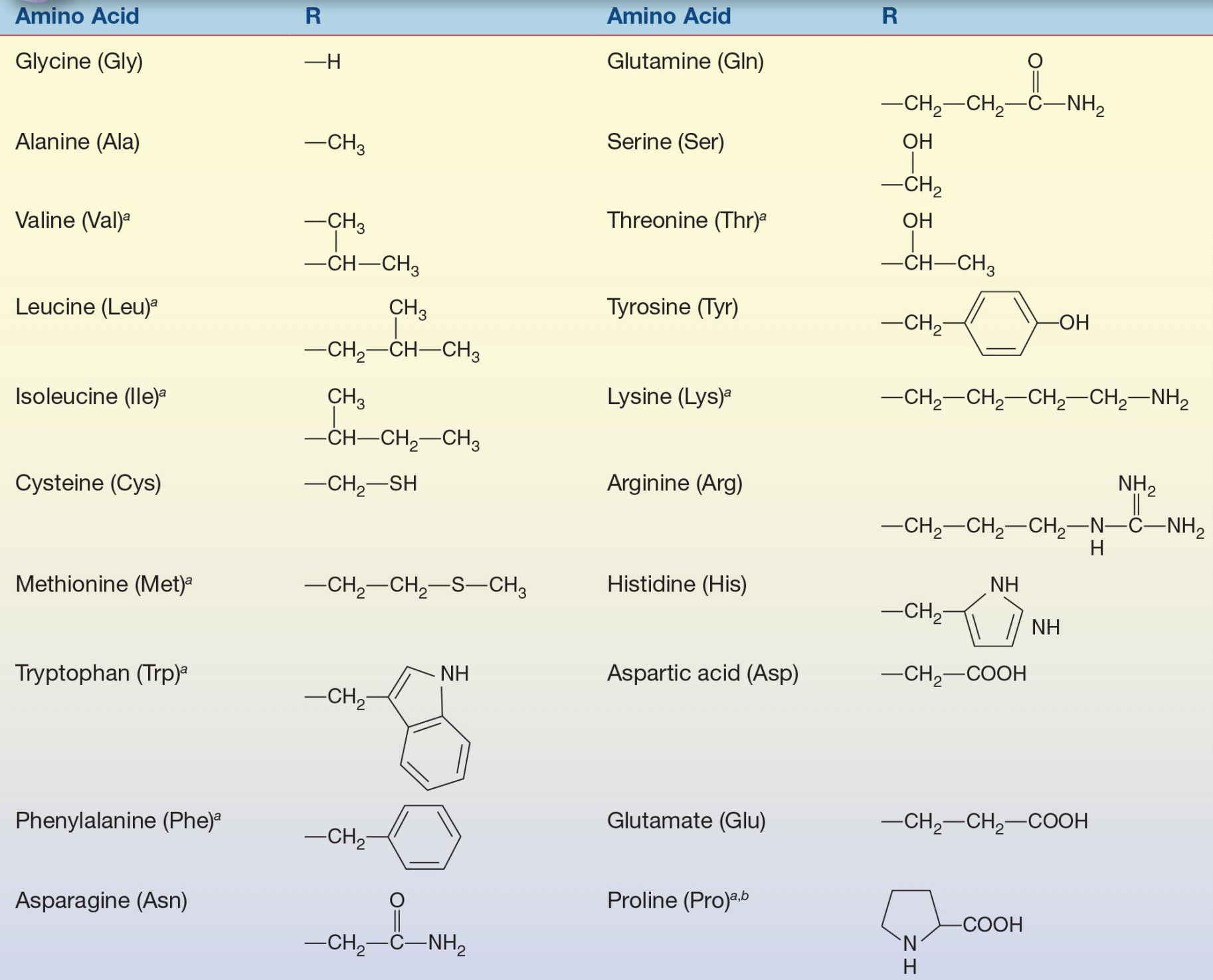
#FAMILY HYDROPHOBIC AMINO ACIDS CODE#
Source code for AlphaFold 2, an algorithm that predicts 3D protein structure with unprecedented accuracy, is now freely available. But researchers Background: Prediction of 3-dimensional protein structures from amino acid sequences represents one of the most important problems in computational structural biology.

RaptorX is a protein structure prediction server developed by Xu group, excelling at predicting 3D structures for protein sequences without close homologs in the Protein Data Bank (PDB). More information may be found in this article's talk page. If you know that your sequences have close homologs in PDB, this server is a good choice. The community-wide Critical Assessment of Structure Prediction (CASP) experiments have been designed to obtain an objective assessment of the state-of-the-art of the field, where SABLE server can be used for prediction of the protein secondary structure, relative solvent accessibility and trans-membrane domains providing state-of-the-art prediction accuracy. This is the PHD/MaxHom server of Rost and Sander.